Key Result
This project concluded that Brassica napus genotype affected soil inorganic nitrogen, extractable sulphur, and total carbon, but had no effect on available P and pH, but genotype effects on soil nutrients were not consistent among three field sites across Saskatchewan. It also determined that soil nutrients varied more strongly over the course of the growing season and it reported that root traits and soil inorganic nitrogen was related to microbial diversity and community composition and crop nitrogen use efficiency.
Project Summary
This project provides added value to the aboveground and root microbiome phenotyping field trials established through funds from the Plant Phenotyping and Imaging Research Centre (P2IRC) developed from the Canada First Research Excellence Fund awarded to the Global Institute for Food Security (GIFS).
The goal of this study was to characterize soil characteristics and crop nutrient uptake to get a better understanding of how these factors influence crop productivity, especially as assessed in breeder trials. This helps to advance breeding efforts to develop profitable crops for producers with minimal environmental impact for all.
The specific objectives being addressed are:
(1) whether crop nutrient uptake profiles and soil nutrient dynamics differ with canola genotype
(2) how dynamic soil properties such as pH and soil carbon that govern nutrient availability interact with genotype to influence nutrient uptake and crop productivity.
Underlying these objectives is also the need to understand within site soil variability (part of the environmental component needed to understand genotype by environment interactions) and how this variability may affect crop productivity and interpretations of phenotypic information across crop genotypes. This soil and plant nutrient data will be related to the rhizosphere, root, and endophyte (shoots and leaves) microbiome data as well as aboveground phenotyping data being collected as part of the P2IRC initiative across a diverse set of canola genotypes.
The field trials were designed to test advanced imaging (e.g., drone-based multispectral), molecular (e.g., microbial metagenomics, synchrotron), and computational tools (e.g., machine learning) to link crop phenotype to genotype, a significant bottleneck limiting the advancement of crop breeding.
Results
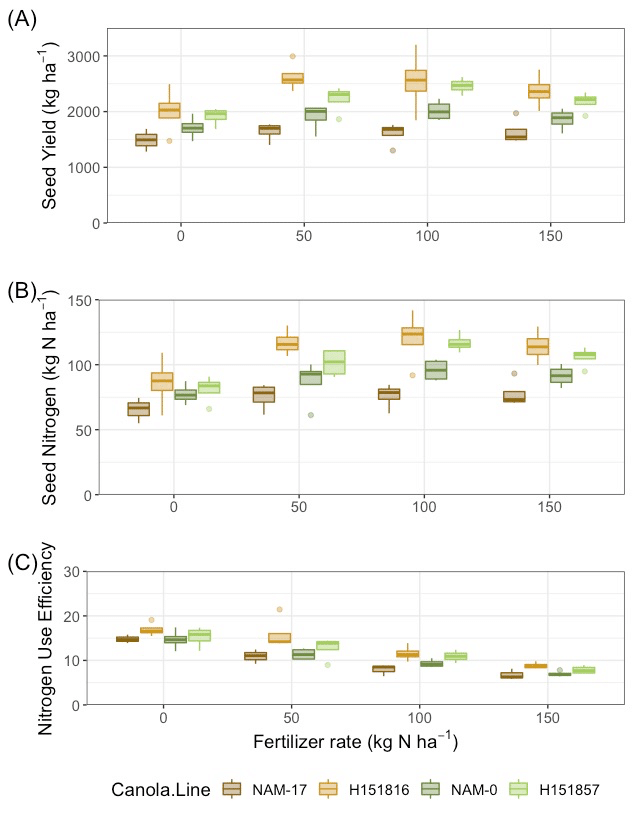
Soil inorganic nitrogen and extractable sulphur were most responsive to differences in B. napus genotype, while available phosphorus and soil pH were unaffected. Variation in soil properties was strongest within the growing season and changes in nutrients were relatively consistent among site-years, with some exceptions. In a more focused investigation of soil nitrogen cycling and nitrogen use efficiency, there was significant relationships between soil microbial community composition, soil inorganic nitrogen, and crop nutrient use efficiency.
Further research in identifying the drivers of genotype-specific differences in soil nutrient cycling and uptake are warranted.





