Key Result
Composed a NAM population of 2500 recombinant inbred lines, in addition to phenotyping parental lines and their progeny, which provided a detailed picture of their genetic composition. This knowledge will be used to analyze the introduction of traits such as drought and heat stress, root-soil interactions, seed dormancy and verticillium resistance into commercially available canola varieties.
Project Summary
Overview
This study aims to broaden the genetic pool available for canola breeding, capturing diversity from all available collections of annual B. napus. The project will also provide the tools for rapidly introducing valuable variation into cultivar development.
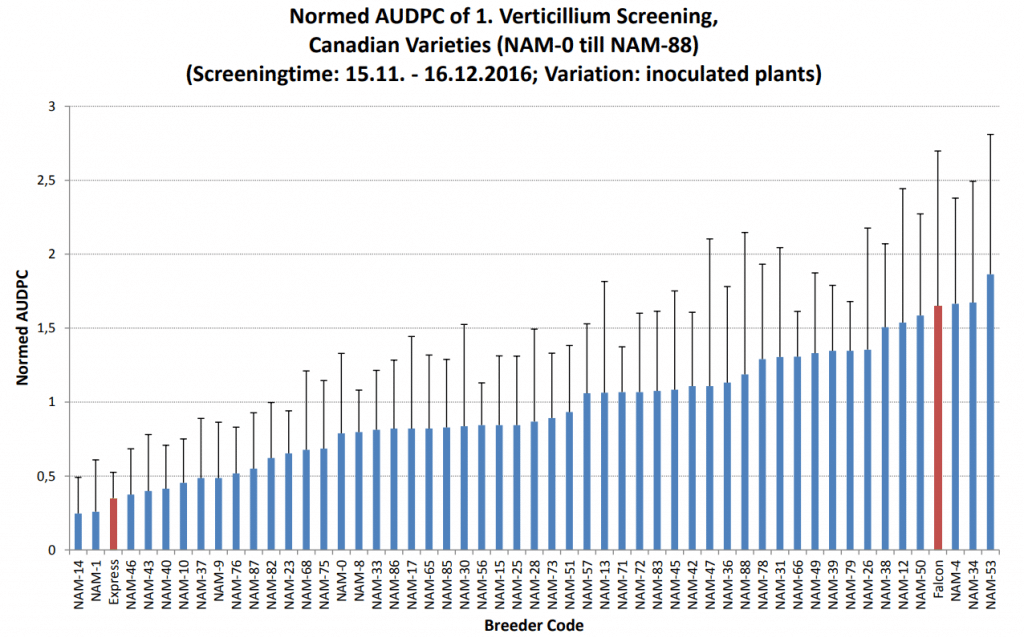
wilt, Falcon and Express are the susceptible and resistant checks, respectively.
Researchers conducted lab, greenhouse and field based studies to phenotype the 50 founder lines and common line, from which the NAM population was developed, for four field seasons under Saskatchewan conditions. In 2015 and 2016, the lines were grown in additional Saskatchewan locations in Melfort, Outlook (irrigated and dryland plots) and Scott, and also at Beaverlodge, Alberta. The molecular and genomics technologies research components, including the NAM population mapping, was conducted at AAFC Saskatoon. The seed multiplication was done by AAFC in collaboration with industry partners.
Results
Researchers developed a structured population of B. napus that is being used for the study of complex traits in both the field and lab environment. The NAM population generated is composed of >2500 recombinant inbred lines derived from 50 diverse crosses. The founding parental lines were extensively phenotyped in the field and lab, highlighting wide ranging variability for agronomically important traits. The whole population was assayed in the field in 2016 providing preliminary data for association of important traits with the underlying genes controlling their expression.
The developed NAM population will be an extremely valuable resource for studying multiple traits relevant to sustainable canola production.





