Key Result
Although minimal impacts of rotation were found on root nutrient availability, root exudate production was affected by continuous canola, resulting in differences in the fungal microbiome associated with canola roots.
Project Summary
Overview
Crop rotation is an important strategy used by producers to maximize soil and plant health. Rotation strategies are complex and involve a variety of biological, economic, and social factors that determine the yearly sequence of crops that are planted. The effects of common rotation strategies on the microbial communities recruited by the plant have not been investigated. We found that the soil and root-associated fungal microbiome responded more strongly to crop rotation compared to the bacterial communities, and identified a fungus, Olpidium brassicae, that was particularly dominant in the canola-only rotations. These findings highlight the importance of considering the effects of crop rotation on the whole plant, including its associated microbial partners, and point to new areas for future research.
Purpose
The objectives of this research included determining the effect of crop rotation on the nutrient fluxes experienced by canola roots during flowering; examining the organic acids produced by canola that shape the composition of the root-associated microbial communities, and characterizing the resulting microbial communities. We aimed to determine the effect of crop rotation on the whole plant, including its microbial partners, providing producers with holistic data on the biological effects of production choices that are informed by a combination of economic, biological, and pragmatic considerations.
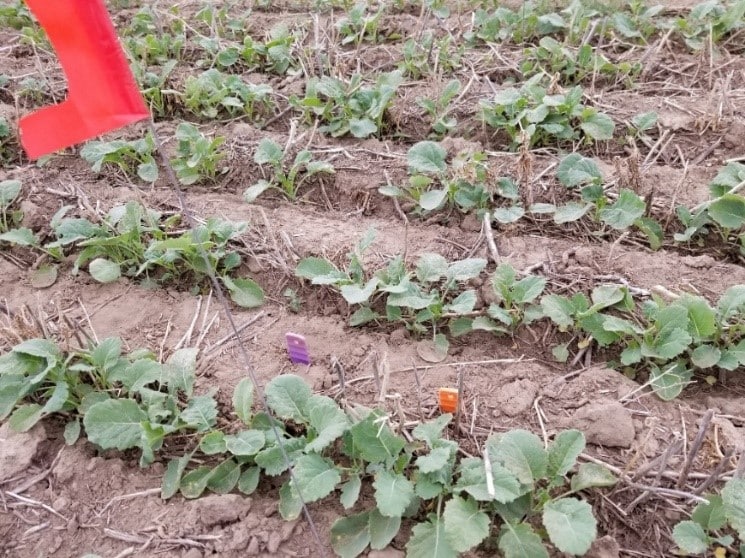

Photo credit: Dumonceaux and Helgason research team
Methodology
This project leveraged the efforts of a long-established crop rotation study that examined the agronomic impacts of crop rotation strategies including canola; canola-wheat; and canola-pea-barley. We examined three locations (Scott, SK; Swift Current, SK; Lacombe, AB) representing a variety of soil types (brown; dark brown; black), and the samples were taken from the canola year of well-established rotations – years 11 and 12 (final years) of the larger study. We used commercially available Plant Root Simulator® probes from WesternAg to examine the flux of nutrients available to canola roots, and ion exchange chromatography to examine the organic acids produced by the canola plants. Bacterial and fungal communities were characterized using DNA sequencing of the soil, rhizosphere and roots. In this way, we examined the long-term impact of crop rotation on soil processes and the canola microbiome to better understand how canola cropping frequency impacts crop performance.
The generation of sequencing data from this project, especially from the second year, was seriously impacted by the lack of laboratory access caused by the covid crisis in early 2020. This caused a delay in the completion of the project, from March 31, 2021 to September 30, 2021. Because the sample collection occurred as planned in 2018/2019 these delays did not affect the quality of the data or interpretation of results. All objectives were completed once laboratory access was re-gained. A technical issue precluded the collection of root exudate data in the first year of the project but this was corrected for the second year.
Results
We found that long-term, short-rotation canola had transient, site-dependent effects on soil nutrient availability. This was likely due to the use of soil test-based fertilization according to normal agronomic practices, which provided a crop responsive prescription for fertilizer addition and ultimately, adequate nutrients for crop growth. In addition, we determined that the crop rotation strategy had a stronger impact on the fungal microbiota at all sites, years, and soil compartments examined compared to the bacterial communities. Crop rotation affected the abundance of fungi associated with both detrimental and beneficial effects on crop production, and this effect was most pronounced in and near canola roots. In particular, continuous canola resulted in a predominance of a fungus identified as Olpidium brassicae, a poorly understood root colonizer with a life cycle that is similar to that of the clubroot pathogen, Plasmodiophora brassicae. This fungus was present in and near the roots at all sites, and was particularly dominant in short-rotation and continuous canola. This provides a direction for future studies.
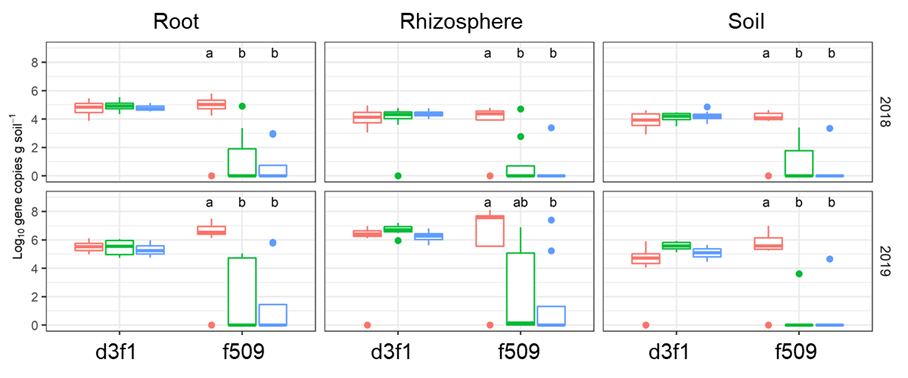
Red bars, continuous canola; green bars, canola-wheat; blue bars, canola-pea-barley. Letters above the bars indicate statistically significant differences in measured abundances. The fungus coded f509 was significantly enriched in continuous canola compared to canola grown in either rotation.
Conclusions
This research examined canola root systems in rotations including canola; canola-wheat; and canola-pea-barley. We found minimal impacts of rotation on root nutrient availability. However, root exudate production was affected by continuous canola, resulting in differences in the fungal microbiome associated with canola roots. A sequence identified with Olpidium brassicae, a fungus with a life cycle similar to that of the clubroot pathogen, dominated the communities, and was especially prevalent in the canola-only rotations.
These findings highlight the importance of considering the impacts of crop rotation strategies on the microbiota associated with canola roots, and demonstrate that the fungal biome is more affected by continuous canola than the bacterial community. Continuous canola resulted in the dominance of a particular fungus, Olpidium brassicae, with unknown effects on canola health and resiliency.
Acknowledgements: We thank Mr. Steve Kessler, Ms. Min Yu, and Ms. Christine Hammond for their contributions and technical expertise.